Same Concept, Two Different Resolutions
In 2021, Oxford researchers published a puzzling finding in Neurology:
They measured "chronic inflammation" in the same group of older adults — once using traditional serum CRP protein levels, and once using a DNA methylation proxy.
Same concept. Same population. The two measurements should have yielded similar conclusions.
They didn't. The DNA methylation proxy was 6.4 times more strongly associated with brain atrophy than the serum protein.
The concept wasn't wrong. The measurement was — or more precisely, the focus was off.

Prism & Focus: The Optics of Gene-Set Representation
Imagine a beam of white light passing through a prism. The white light is a biomedical concept — say, "chronic inflammation." The prism refracts it into multiple colored beams, each representing a different measurement modality: serum protein, transcriptome, DNA methylation, metabolites...
These beams land on different focal planes. Only one beam forms a sharp, clear image on the correct plane — that focal point is the gene set. The sharpness of the image is the strength of association with disease.
Most concepts are currently out of focus. We've been using the wrong measurement modality, seeing blurry images, and mistakenly blaming the concept.
EvoSika's job is to systematically find the right prism and the right focal length for every biomedical concept.
And methylation — with its "long exposure" nature — is naturally suited to focus on slow-accumulating chronic processes.
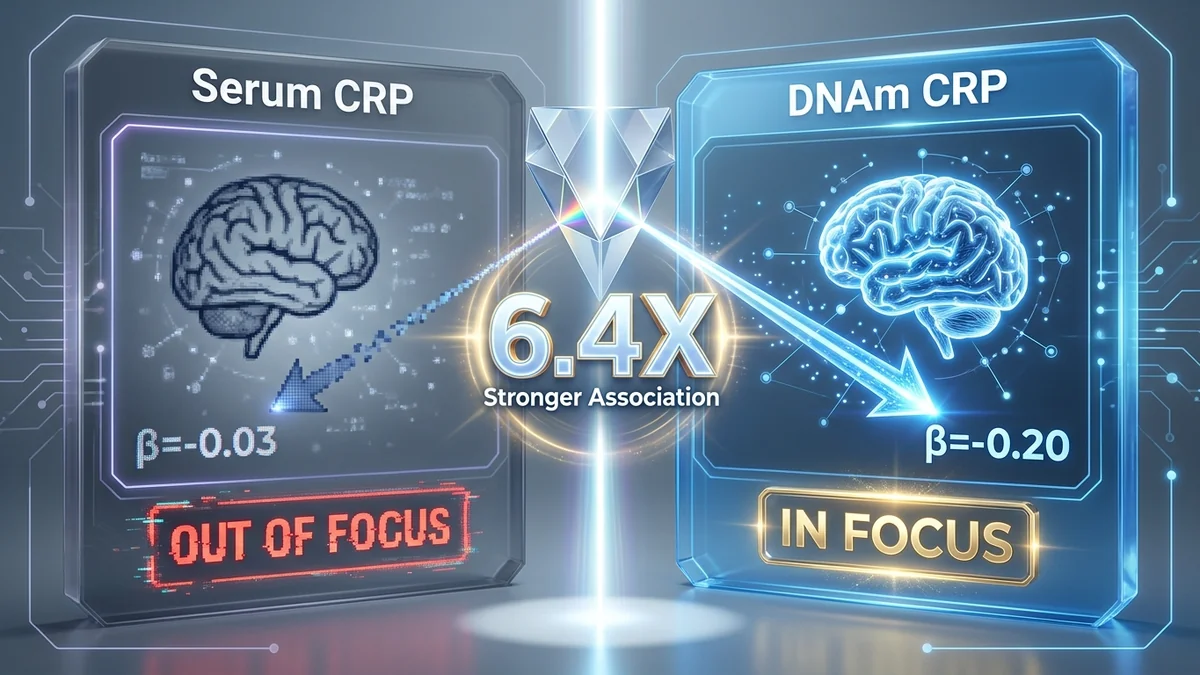

The Chronic Inflammation Focus Test
This is the empirical study published by Conole et al. in Neurology (2021). We reframe it as a "focus experiment":
| Serum CRP (Protein) | DNAm CRP (Methylation) | |
|---|---|---|
| What it measures | Instantaneous inflammatory protein concentration | Long-term inflammatory exposure recorded in methylation marks |
| Stability | Low (up to 100-fold within-day fluctuation) | High (month-to-year scale changes) |
| Association with total brain volume | β ≈ -0.03 | β = -0.197 (p = 8.42×10⁻⁶) |
| Association with gray matter volume | Very weak, non-significant | β = -0.200 (p = 1.66×10⁻⁵) |
| Focus status | ❌ Out of focus | ✅ In focus (6.4× stronger) |
Why?
Because CRP protein in blood is a snapshot — an infection, a sleepless night, or a high-fat meal can make it spike dramatically. You might be measuring last night's poor sleep, not true "chronic inflammation."
DNA methylation, by contrast, is a long exposure — it changes slowly and integrates years of inflammatory exposure. The signal is amplified, the noise averaged out.
Same concept. Different measurement modality. 6.4× stronger association. That's the power of getting the focus right.
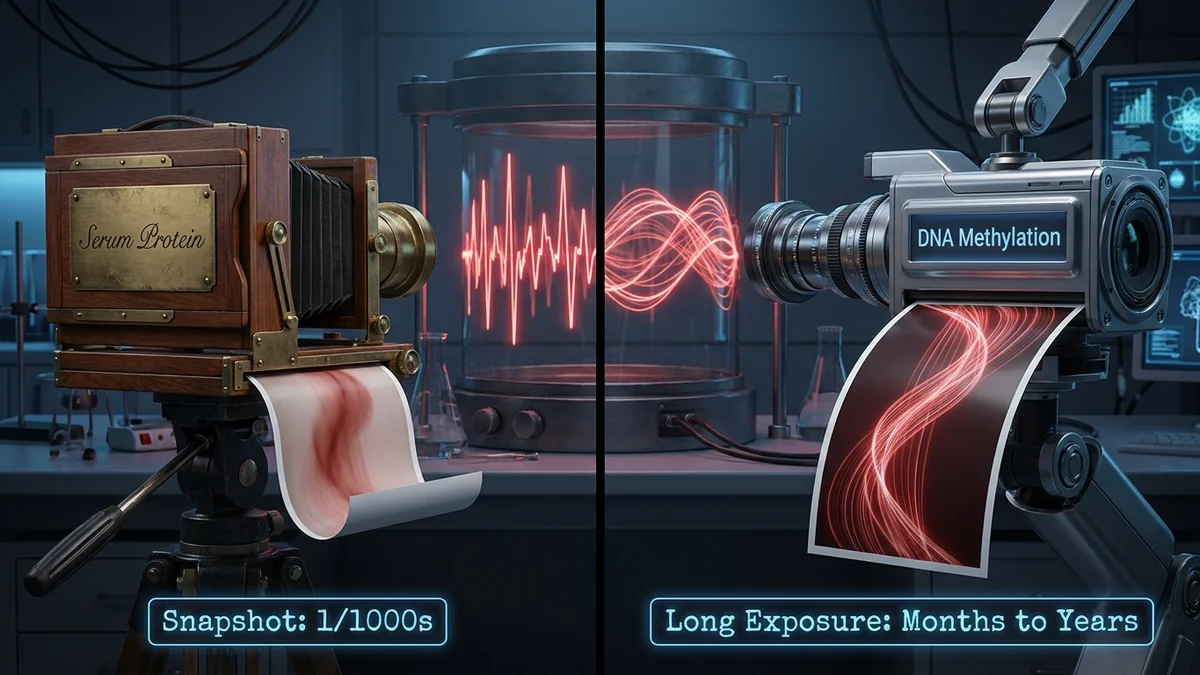

EvoSika's Four-Dimensional Benchmark: Focusing, Systematized
Conole's paper demonstrated a brilliant single focus — but it was one concept, one researcher's intuition.
EvoSika systematizes the process. We break "focus" into four automatically computable dimensions:
Causal Emergence
Is the gene set's signal strong enough for the concept to "emerge"? This is exactly what the CE Index measures — serum CRP's CE Index is low (signal drowned in noise), while DNAm CRP's CE Index is high (signal emerges).
Parsimony
How many genes are actually needed? Redundant genes defocus the image.
Cross-Disease Generalization
Does this gene set stay in focus across multiple diseases, or only in one?
Intervention Efficacy
Can it guide clinical decisions? Is the image sharp enough to operate on?
Four dimensions. One clear score. From "guessing which measurement to use" to "systematic focus engineering" — this is EvoSika.
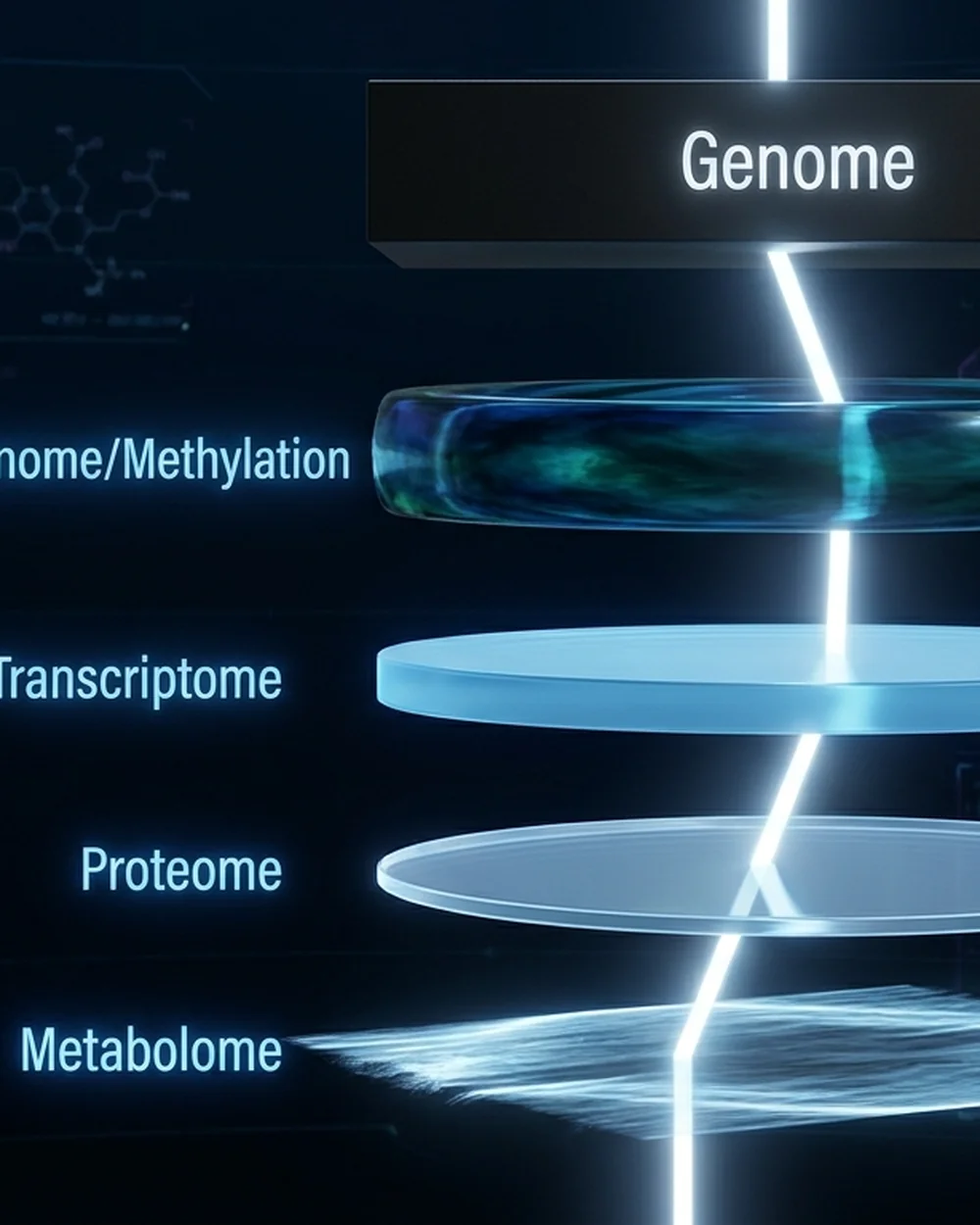
The Multi-Omics Exposure Timeline: Different Concepts Need Different Shutter Speeds
The CRP focus experiment hints at a deeper principle — one that points to a fundamental question precision medicine has not yet systematically answered:
What is the optimal "shutter speed" for a given concept?
Let's reframe each omics layer in terms of exposure time:
| Omics Layer | Rate of Change | Shutter Speed Equivalent | Best for Focusing On | Not Good For |
|---|---|---|---|---|
| Metabolome | Seconds to hours | 1/1000s | Acute drug response, exercise stress | Chronic aging, long-term immune memory |
| Proteome | Hours to days | 1/100s | Short-term metabolic states, acute phase | Cumulative organ damage, chronic low-grade inflammation |
| Transcriptome | Days to weeks | 1/10s | Cell state switching, circadian rhythm | Multi-decade degenerative processes |
| Epigenome | Months to years | Long exposure | Aging accumulation, developmental programming, long-term environmental exposure | Immediate physiological response |
| Genome | Lifetime-stable | The film itself | Developmental framework, genetic risk | Dynamic processes (unmeasurable) |
This isn't about methylation being "better." It's about this: for "chronic" concepts, you need slow measurements to get a sharp picture; for "acute" concepts, fast measurements are what capture the moment.
The problem is that most aging and chronic disease concepts are "slow concepts" — yet most traditional biomarkers use "fast measurements."
EvoSika finds the optimal exposure time for every concept. This is the ultimate question that gene-set representation asks: not just "what to measure," but "how long to measure for."
Every Concept Is a Prism
If we push the prism-focus logic one step further, a deeper recursive structure emerges:
A concept is itself a prism.
The same biomedical reality — say, "aging" — can be refracted through different conceptual prisms: chronic inflammation is one, mitochondrial dysfunction another, epigenetic alteration a third. Each prism casts different beams, and each beam demands a different focal length.
Even the question of which concept to use for describing a disease needs to be brought into focus.
EvoSika lets concept compete against concept — their gene sets face off in the same benchmark arena. The fittest survive on the leaderboard; the flawed are eliminated; two excellent concepts spontaneously merge into a better one. Just as light is refocused after passing through a prism, concepts are redefined after passing through evaluation.
This is EvoSika's deepest conviction: knowledge itself is a form of life, and evolution is its only rule.
Further reading: Philosophy & Roadmap →