Benchmark Framework
EvoSika employs a four-layer evaluation architecture, from Causal Emergence to Intervention Efficacy, comprehensively assessing the scientific and clinical value of gene sets.
Module 1 — Causal Emergence
Evaluates whether a concept exhibits stronger disease associations than individual genes. Quantifies the overall emergence effect through the Causal Emergence Index (CE Index), verifying whether the gene set truly achieves a "1+1>2" emergence effect.
Core Metrics: CE Index (Causal Emergence Index), AUC (Disease Classification Accuracy)
Module 2 — Parsimony
Evaluates whether the gene set can achieve equivalent disease classification with the fewest variables. Automatically selects key genes through ElasticNet regularization (L1+L2) and assesses the parsimony score.
Core Metrics: Parsimony Score, Number of Non-Zero Coefficients
Module 3a — Pan-Disease Explanatory Power
Evaluates whether the gene set possesses universal explanatory power across multiple diseases. Validates generalization capability through cross-disease datasets (breast cancer, colorectal cancer, depression, etc.).
Core Metrics: Universal Disease Score, Mean Cross-Disease AUC
Module 3b — Intervention Efficacy
Evaluates whether the gene set responds to effective interventions. Validates intervention responsiveness using real GEO clinical trial data (folic acid, flavanols, metformin, etc.).
Core Metrics: Universal Intervention Score, Pre- vs. Post-Intervention Significance
Evolution Mechanisms
EvoSika's evolution engine draws on the natural wisdom of sika deer antler shedding and regeneration to achieve continuous iterative optimization of theories.
Fitness exceeds threshold (≥0.5), new concept is directly added to the gene set library
Highly similar to an existing Agent (Jaccard≥0.8), merged into a new version
Clear internal substructures detected within the concept, splitting into multiple sub-Agents
Outperforms an existing similar Agent (Jaccard 0.3-0.8), replacing the old version
Insufficient fitness, concept is eliminated
Distributed Computing Toolkit
Download the EvoSika computing package, run benchmarking tasks locally, and submit results to the leaderboard.
Supports Windows / macOS / Linux
The computing package includes: benchmarking scripts + reference datasets + SHA-256 gene set fingerprinting
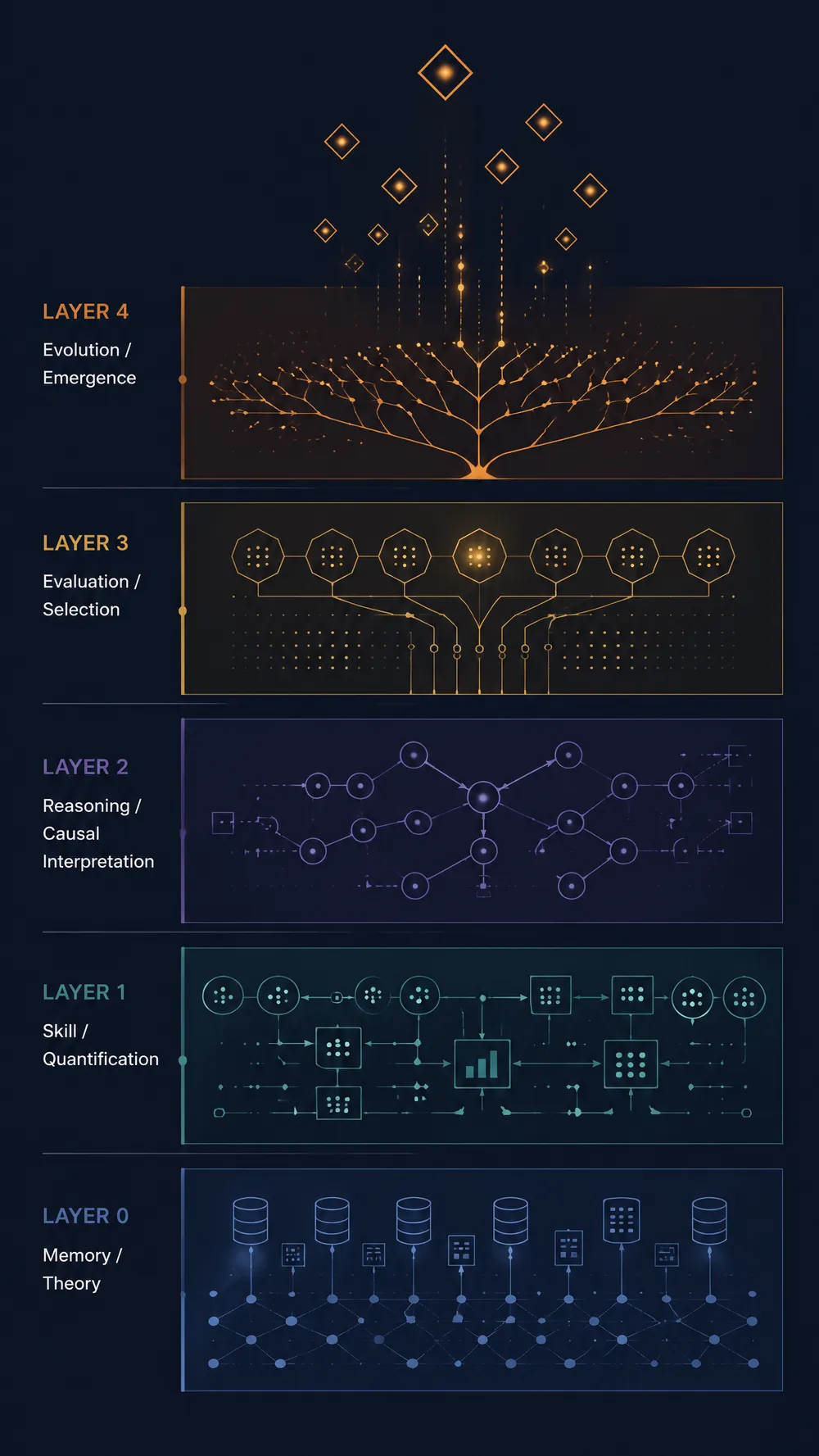
Four-Dimensional Evaluation Framework

Layer 1: Causal Emergence
Is this concept truly causally related to disease? Quantifying macro-emergence effects through the Causal Emergence Index (CE Index) — Hallmark-level features can be up to 9.7 orders of magnitude stronger than individual genes.
Layer 2: Parsimony
Does this gene set representation achieve equivalent predictive accuracy with the fewest variables? Evaluating parsimony scores through LASSO/ElasticNet.
Layer 3: Pan-Disease Explanatory Power
Does this concept have universal explanatory power across multiple diseases? Validated across 10 age-related disease datasets.
Layer 4: Intervention Efficacy
Can this concept distinguish effective interventions from ineffective ones? Using real GEO clinical trial data to verify significant changes before and after intervention and therapeutic mediation effects.
Three Steps to Participate
Define Your Concept
Enter a biological concept name (e.g., "Mitochondrial Dysfunction", "Qi Deficiency") and submit the gene set you believe best characterizes it. The system automatically registers it as a standardized Hallmark Agent.
AI Automated Evaluation
Your Agent enters the Hallmarks Engineering Testbed evaluation queue. The offline evaluation engine automatically performs four-dimensional evaluation (causal emergence, parsimony, pan-disease explanatory power, intervention efficacy) on public datasets. Estimated completion within 12 hours.
View Rankings & Evolution
Upon evaluation completion, your Agent appears on the public leaderboard. You can see its precise ranking on each disease leaderboard, each evaluation dimension sub-leaderboard, and the cross-disease overall leaderboard. Winners are retained, flawed ones eliminated, and two excellent concepts can spontaneously merge.